Home
New at SPC
About the Center:
Overview
Center Structure
People
Center Research
Tech Modules
Pilot Projects
BWG
HTPF WG
Macrophage WG
Administrative Core
Resources
Protocols
Training
Publications
Contact the Center
|
| |
|
| |
PROTEOMICS TOOLS
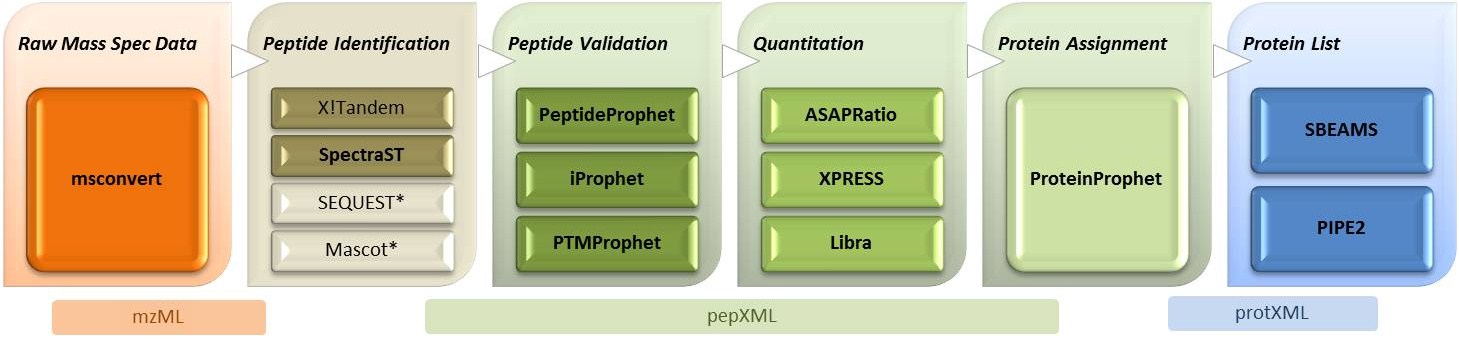
The Trans-Proteomic
Pipeline (TPP)
includes all of the steps of the ISB MS/MS analysis pipeline,
after the database search. Most users will only need to download the
TPP distrubtion.
Download the TPP Installer for Windows
The individual tools that make up the TPP, as well as a few which are
not included in the TPP distrubution, are categorized below.
Please see the SPC Tools Wiki for more information, including download information.
- Input Processing / Data Handling
- mzXML - standard format for mass-spectrometry data
- ReAdW - Thermo Xcalibur-to-mzXML converter
- massWolf - Waters MassLynx raw-to-mzXML converter
- mzWiff - ABI/MDS Sciex Analyst raw-to-mzXML converter
- mzBruker - Bruker raw-to-mzXML converter
- RAP, JRAP, RAMP - mzXML data parsers
- validateXML - mzXML validation script
- MzXML2Search - mzXML to SEQUEST dta, MASCOT generic
and Micromass pkl converter
- mzXMLViewer - viewer for displaying mzXML data
- InsilicosViewer - viewer for displaying mzXML data
- readmzXML - mzXML parser based on RAMP
- Database and Spectral Library Search
- SpectraSTTM - spectral library search engine provided by TPP
- Comet - open-source database search engine provided with TPP
- X!Tandem - open-source database search engine provided with TPP
- Probability Assignment and Validation
- PeptideProphetTM - validation of PSMs made by tandem mass spectrometry (MS/MS) and database searching;
probabilities are assigned to the peptide identifications made by programs like SpectraST, Comet or X!Tandem
- iProphetTM - validation of distinct peptide sequences; can also combine search results of multiple search engines
- ProteinProphetTM - statistical model for
validation of peptide identifications at the protein level
- Protein Quantification
- XPRESS - software to calculate the relative abundance of proteins in the sample
- ASAPRatio - Automated Statistical Analysis on Protein Ratio
- Libra - Four channel quantification software
- Protein ID Curation
- QualScore-- analyze unassigned but high quality spectra.
- PepXMLViewer - peptide dataset interaction
- Pep3D - viewer for LC-MS and LC-MS/MS results
- Out2Summary - converter of SEQUEST and TurboSEQUEST *.out files for TPP processing
into a single HTML-SUMMARY file ready for use with INTERACT
- Protein Crosslinking
- Miscellaneous
- LC-MS Analysis
- SpecArray - tool for analyzing and comparing LC-MS runs
- SuperHirn - label-free quantitative analysis.
- msBID - alternate tool for MS Level 1 analysis.
- Interpretation
- SBEAMS - Systems Biology Experiment Analysis Management System
- Cytoscape - platform for visualizing and integrating molecular interaction networks
- Prequips: Prequips is an interactive software environment for integration, visualization and analysis of LC-MS/MS data.
|
| |
|


