Software:TPP
From SPCTools
The Trans-Proteomic Pipeline (TPP) is a collection of integrated tools for MS/MS proteomics, developed at the SPC.
Contents |
Overview
Authors and Contributors
See TPP:Authors and Contributors
Getting the software
The latest TPP version is 7.3.0, released April 2025.
For support both during and after installation, you are encouraged to consult the SPC Tools newsgroups:
|
Installing on a Windows System
The Windows version of TPP is provided as a pre-built binary self-installing executable which can be downloaded from the SourceForge site here. There are also detailed instructions on how to install the latest version of TPP in the TPP 5.2.0 Windows installation guide (instructions also apply to 6.x and 7.x versions).
Source Code Installation (For Linux and other systems)
The latest source code package can be found here, on the Sashimi project site on SourceForge. This is gzipped tar archive that can be unpacked using tools such as tar, 7-zip, or WinZip. Details on how to compile TPP can be found in the README and INSTALL_LINUX files found in the top level directory of the package. There is also a series of installation guides, which are more detailed and contain information specific to certain distributions: Linux Installation Guides.
Advanced Topic: building the TPP from source on Windows
NOTE: These advanced topics are for developers. Windows users can download and run the TPP Windows installer, which includes everything needed to run the TPP.
- TPP:Building Windows-native binaries with Mingw
- TPP:Building Windows-native binaries with Visual Studio 2005
Mac OSX
Please Note: TPP 7.1.0 is the latest officially supported version on Mac OSX. Please download the MacOS package
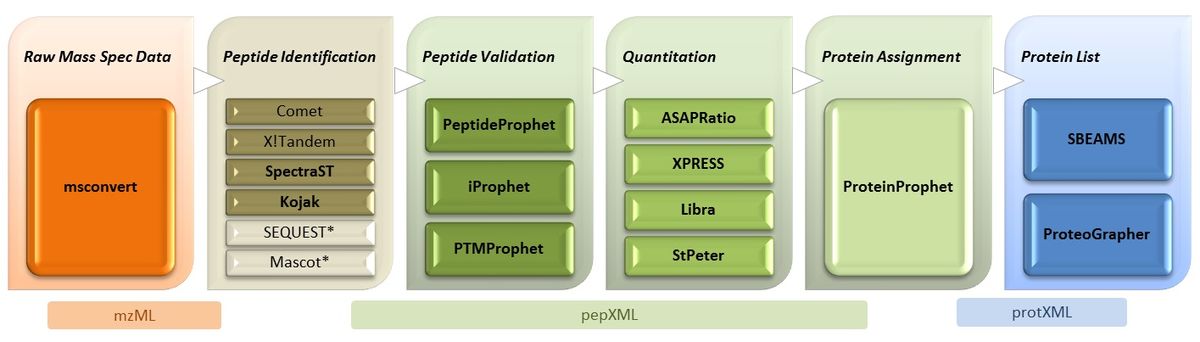
Software contained in the TPP
Probability Assignment and Validation
PeptideProphet: Statistical validation of PSMs using search engine results.
iProphet: Distinct peptide sequence validation, using PeptideProphet results; can also combine the results of multiple search engines.
ProteinProphet: Protein identification and validation, using PeptideProphet OR iProphet results.
Mayu: Decoy-estimated FDRs (false discovery rate) for PeptideProphet results.
Protein Quantification
XPRESS: Calculation of relative abundances of peptides and proteins from isotopically labeled MS/MS samples.
ASAPRatio: Automated Statistical Analysis on Protein Ratio; an alternative to XPRESS.
Libra: Quantification of isobarically-labeled samples (e.g. iTraq, TMT, etc) for any number of channels
StPeter: Label-free Quantification of Shotgun-MS using Normalized Spectral Indexes, Spectral Counts, or Normalized Spectral Abundance Factors.
Graphical User Interface (GUI)
Petunia: Petunia is the name of the TPP's web-based GUI, which presents the tools in an organized and logical manner for those who do not wish to use the command-line.
Spectral Library Building and Searching
SpectraST: Searches spectral libraries (including publicly available ones from NIST and GPM) to identify peptide MS/MS spectra. Builds spectral libraries from sequence search results.
Protein ID Curation
Out2Summary - converter of SEQUEST and TurboSEQUEST *.out files into a single HTML-SUMMARY file ready for use with INTERACT
Pep3D: Viewer for LC-MS and LC-MS/MS results.
Input Processing: mzXML Tools
readmzXML: mzML/mzXML parser based on RAMP
MzXML2Search: Converts mzXML files to SEQUEST dta, MASCOT mgf, and Micromass pkl files
mzStar: SCIEX/ABI Analyst format to mzXML converter
ReAdW: ThermoFinnigan Xcalibur format to mzXML converter
MZIdentML: Handling MZIdentml
DidIScanThat: Interrogating MS/MS spectra of mzML files.
RAMP: mzXML data parser
Input Processing: Search-Engine to pepXML converters
- Sequest results: Out2XML
- Mascot results: Mascot2XML
- Tandem results: Tandem2XML
Database Search Tools
- X!Tandem: Open source peptide search engine from GPM
- Comet: Open source peptide search engine from Comet-MS
Working with supported search engines
The TPP currently supports Sequest, Mascot, ProbID, X!Tandem, Comet, SpectraST, MSGF+, Inspect, MyriMatch, and Phenyx. Please see the supported search engines page for more information.
Example Data Analysis
- Example Data Analysis
- Try our TPP Tutorial
- You can also see the older TPP Tutorial v2 and TPP Tutorial v1
TPP and Related Software Tools
This page describes the (sometimes confusing!) relationship between the many software projects that integrate with the TPP, either as compatible search engines, or as projects that repackage and redistribute the TPP itself.
Additional help
FAQ
Newsgroups
subscriptions highly recommended for SPC Tools users
spctools-discuss
spctools-discuss discussion group: very active, daily discussions ranging from installation to data processing. All users, new and experienced, encouraged to participate.
spctools-announce
spctools-announce discussion group: infrequent, important notifications of updates to our software
TPP Demo and Tutorial
Try or end-to-end analysis of Orbitrap SILAC data in our TPP Tutorial.
Developer Documentation
- Developer Documentation
- Boost CPP libraries
- Debugging:Understanding XSL-Generating Code in Perl
- Debugging:Working with boost::shared ptr