Software:TPP
From SPCTools
The Trans-Proteomic Pipeline (TPP) is a collection of integrated tools for MS/MS proteomics, developed at the SPC.
Software contained in the TPP:
Contents |
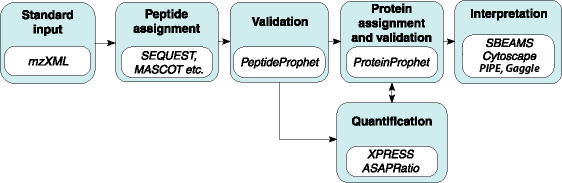
Probability Assignment and Validation
PeptideProphet: Statistical validation of spectra-to-peptide sequence, using search engine results.
ProteinProphet: Protein identification and validation, using PeptideProphet results.
Protein Quantification
XPRESS: Calculation of relative abundance of proteins from MS/MS data.
ASAPRatio: Automated Statistical Analysis on Protein Ratio.
Libra: Four channel quantification software.
Graphical User Interface (GUI)
Petunia: Petunia is the name of the TPP's web-based GUI, which presents the tools in an organized and logical manner for those who do not wish to use the command-line.
Protein ID Curation
Out2Summary - converter of SEQUEST and TurboSEQUEST *.out files into a single HTML-SUMMARY file ready for use with INTERACT
Pep3D: Viewer for LC-MS and LC-MS/MS results.
Input Processing: mzXML Tools
readmzXML: mzXML parser based on RAMP
MsXML2Other: mzXML to SEQUEST dta, MASCOT generic and Micromass pkl converter
mzStar: SCIEX/ABI Analyst format to mzXML converter
ReAdW: ThermoFinnigan Xcalibur format to mzXML converter
RAMP: mzXML data parser