TPP Tandem search
From SPCTools
Performing XTandem search using the Trans Proteomic Pipeline (TPP) Tutorial
TPP V4.0.2, 2008. Note: Screenshots may vary from the TPP build you are using because the application is in development.
This document was originally assembled by Lik Wee Lee of ISB. (work in progress)
Contents |
Introduction
This tutorial will cover the application of the Trans Proteomic Pipeline (TPP) to do a database search of acquired MS/MS spectra using X!Tandem. The data used in this tutorial comes from this TPP Tutorial. The TPP Tutorial begins with passing the results from a SEQUEST search through TPP in order to statistically evaluate and quantify the proteins identified. This tutorial on the other hand will cover omitted steps: using TPP to perform a database search. The search engine X!Tandem will be used in this tutorial as it has two advantages: 1) no commercial license is needed to use the software 2) distributed with the TPP installation.
Systems Requirements
This tutorial requires that TPP be installed. TPP is distributed for both Linux and Windows platform, however this tutorial will focus on the Window platform. Since the graphical interface of TPP ( Petunia) is through a web browser, either Internet Explorer or Firefox is required. Although the search engine X!Tandem is required, no separate installation is needed as it will be installed as part of the TPP installation. The TPP installer can be found at sourceforge. The current version (at the time this was written) is 4.0.2. Download the installer: TPP_Setup_v4_0_JETSTREAM_rev_2.exe.
The mass spectrometer data is provided in mzXML format and can be downloaded via www.insilicos.com/spctools/data/tutorial_wiki.exe.
mzXML is a instrument independent data format and the rationale is to provide a standard format that can used by various data analysis software like TPP. Various proprietary and raw file format from mass spectrometer vendors can be converted to mzXML.
Getting Started
Donwloading and installing the TPP
Information on installing and downloading the Windows distribution of TPP can be found at: sourceforge
Setting up an account
The TPP GUI comes with one user account. This account has ‘guest’ as both the user name and password. It is useful to create another account for this tutorial.
Open the DOS shell by selecting Run under the Start menue and typing cmd. In the shell type:
cd c:\Inetpub\tpp-bin\users\
mkdir tandem
cd tandem
crypt isbTPPspc TPP > .password
and
chmod -R 777 C:\Inetpub\tpp-bin\users\tandem
You have just created the password ‘TPP’ for the user ‘tandem.’
In order to add a different username, create a tpp-bin/users/NEWUSER/ directory and run crypt isbTPPspc NEWPASSWORD > .password from this directory. In these examples "isbTPPspc" is the crypt key. This can be changed.
Getting the tutorial data
Unzip the tutorial data into the directory:
C:\Inetpub\wwwroot\ISB\data\Tandem_Tutorial
This directory will now contain 6 mzXML file: raft4041.mzXML, raft4243.mzXML, ... , raft5051.mzXML and a sequest.params file.
There is also a directory named dbase, with subdirectory IPI containing the database file ipi.HUMAN.fasta.v2.31.
Running X!Tandem
Opening the GUI
The TPP pipeline GUI can be opened by clicking on the ‘TPP Web Tools’ shortcut that was created on your desktop during installation or by selecting “TPP Web Tools” under “TPP” in the Windows start menu. Alternatively, you can click on the following link or open your favorite web browser and paste this link into the navigation bar:
http://localhost/tpp-bin/tpp_gui.pl
Login as ‘tandem’ and use ‘TPP’ as the password.
This tutorial is written from the point of view of a researcher viewing data on the computer where the TPP tools are running.
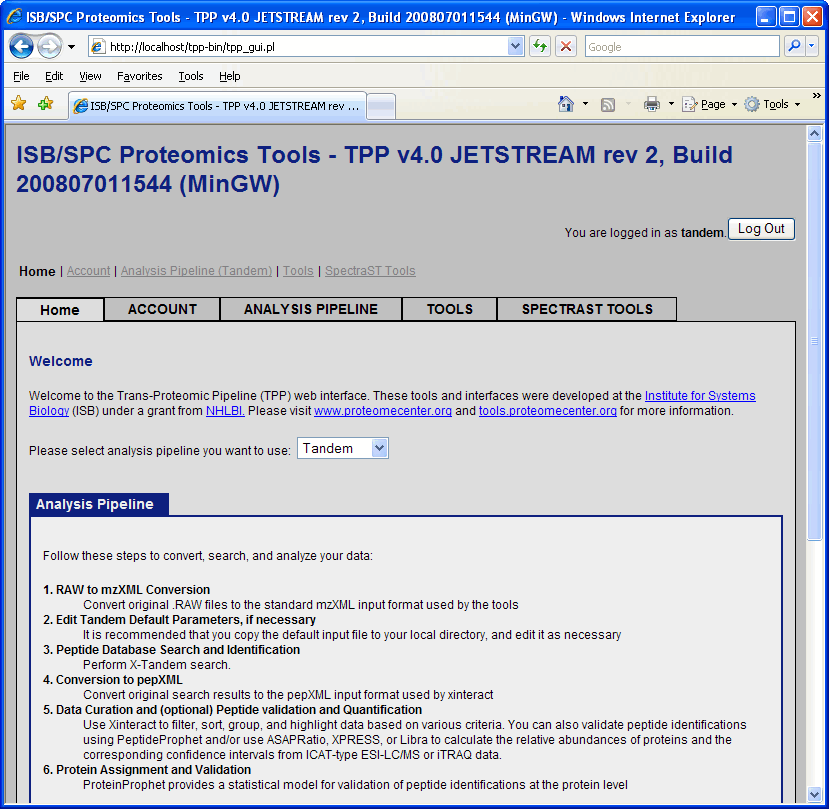
At this point you will be in the “Home” tab of the proteomics pipeline GUI. The Home tab contains information about TPP and the structure of the GUI, along with a pull down menu that lets you choose between SEQUEST, Mascot, Tandem or SpectaST. The default is SEQUEST. Click and change this to Tandem.
Database search using X!Tandem
Click on “Analysis Pipeline”. This will display six tabs which activate different parts of the pipeline. The first tab is Home, which contains information about the TPP. The second and third tab is used to convert raw data from different spectrometers into mzXML and mzML, and the fourth tab is used to search the ms data which is our immediate goal. Click on the “Database Search” tab.
Configuring X!Tandem parameters
The ms data has already been searched using Sequest. We would like to perform a similar search using X!Tandem through the TPP interface. To do this, we should take into account the relevant static and variable modifications. The following lines in the sequest.params tells us these:
diff_search_options = 8.0 C 0.0 x 16.0 M add_C_Cysteine = 442.2000 ; added to C - avg. 103.1388, mono. 103.00919
Note that the sample data were ICAT labeled (Old-ICAT, light tag = d0 442, heavy tag = d8 450). This modifies adds to 442.2 Da to cysteine. Due to the presence of the isotope deuterium, the heavy tag would have an additional 8 Da over the light tag. This accounts for the 8.0 C in the variable modification parameter of SEQUEST (diff_search_options). The 16.0 M accounts for the presence of oxidized methionine.
Our X!Tandem parameters would contain the corresponding line:
<note type="input" label="residue, modification mass">442.2000@C</note> <note type="input" label="residue, potential modification mass">8.0@C,16.0@M</note>
Other pages containing useful information
- Installing TPP
- A list of software that TPP integrates
- converters to mzXML (an open data format for storage and exchange of mass spectroscopy data)
- TPP related tools
- Using Petunia (which is the graphical interface of TPP)
- TPP:X!Tandem and the TPP
- Example data analysis in TPP
- A very comprehensive tutorial on using TPP to analyzed results from a database search